
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|
|
Due to their distinctness in morphology and life history, Giant-Skippers have been treated by many authors as a Family (Megathymidae) within the Superfamily Hesperoidea (Skippers). Thus Hesperoidea were divided into two families: Giant-Skippers (Megathymidae) and Skippers (Hesperiidae). Even when treated as a Subfamily, it was frequently assumed that their position in the tree of Skippers was basal, i.e. Giant-Skippers were positioned closest to moths and "deeper" than any other branch of skippers.
While most authors acknowledge some similarities between Giant-Skippers and Grass-Skippers (Hesperiinae), such as resting posture, larval feeding on monocots, mutual location of M1, M2 and M3 veins on the forewing, and distribution of the 1st instar larval setae (to name a few features), the differences between Giant- and Grass-Skippers seemed quite significant to justify the placement of Giant-Skippers at the root of the Skipper tree. At one point I tried to rationalize this suggested basal placement of Giant-Skippers and similarities to Grass-Skippers by assuming that the Spread-Wing Skippers ("Pyrginae') might have been derived from Grass skippers.
Here we present and analyze DNA sequences relevant to the placement of Giant-Skippers in the tree. In addition to Giant-Skippers, 2 other skipper groups were chosen: the Awls (Subfamily Coeliadinae) and Grass-Skippers (Subfamily Hesperiinae). Where possible, DNA sequences of type-genera from these groups were taken, i.e. Megathymus, Coeliades and Hesperia. As an outgroup (taxon that is more basal in the tree than the most basal taxon in the group), Macrosoma (Hedylidae) was chosen. We assume that due to its moth-like appearance, Macrosoma branches off closer to the root than any of the three Skippers, and thus is an outgroup. For an outgroup to be most effective for the analysis, it should be as close as possible to the group. There is good evidence, both morphological and molecular, that Hedylidae are the sister group of butterflies and skippers.
With Macrosoma being an outgroup, and thus being positioned at the root, only three different trees of these 4 taxa are possible. Tree 1 positions Coeliades at the base. Tree 2: positions Megathymus at the base. Tree 3: positions Hesperia at the base. No other options exist.
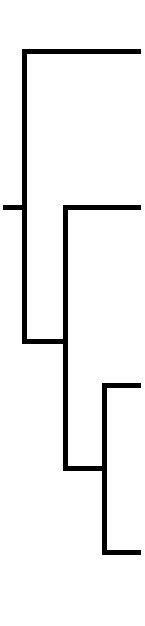 |
 |
| 1. Macrosoma tripulata | |
 |
|
| 2. Coeliades forestan | |
 |
|
| 3. Megathymus streckeri | |
 |
|
| 4. Hesperia leonardus |
Tree 1: Coeliades forestan (Coeliadinae) is a basal group
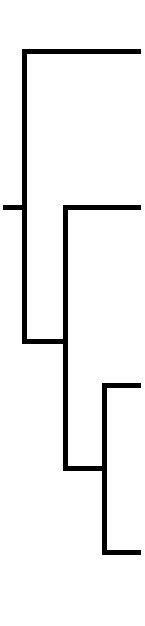 |
 |
| 1. Macrosoma tripulata | |
 |
|
| 3. Megathymus streckeri | |
 |
|
| 2. Coeliades forestan | |
 |
|
| 4. Hesperia leonardus |
Tree 2: Megathymus streckeri (Megathymin?) is basal
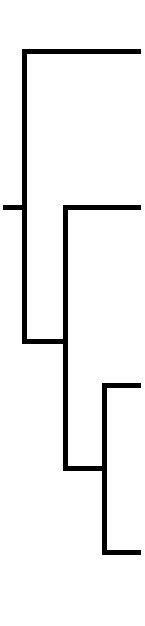 |
 |
| 1. Macrosoma tripulata | |
 |
|
| 4. Hesperia leonardus | |
 |
|
| 3. Megathymus streckeri | |
 |
|
| 2. Coeliades forestan |
Tree 3: Hesperia leonardus (Hesperiinae) is a basal group
All data available to us show support only for the tree 1, in agreement with recent studies (Warren et al. 2008, Warren et al. 2009), while tree 2 is the least parsimonious option.
Methods: All sequences used in this study were obtained from the GenBank database as submitted by Warren et al. (2008) and Weller & Pashley (1995) . Sequences were aligned using the MUSCLE server at EBI. The trees were reconstructed using 4 phylogenetic methods as implemented by the phylogeny.fr server with default parameters, and visualized with ATV and TreeView. Tutorial about how to perform these procedure is available from here.
Data: Partial DNA sequences of two nuclear (Elongation factor-1 α [EF1-a] and wingless [wg]) and two mitochondrial (cytochrome c oxidase subunit I [COI] and NADH dehydrogenase subunit 1 [ND1]) genes, their concatenated alignment, separate alignments (EF1, WG COI, ND1), and trees built from concatenated alignment by TNT, BioNJ, PhyML, MrBayes can be downloaded as text files using the links in this sentence.
Discussion: Currently, there are 4 major classes of methods to reconstruct trees. These methods are based on quite different principles, except that the general idea of Occam's razor (minimize the number of probable DNA changes) in applied everywhere. It is hardly possible to imagine a phylogenetic method that is not based on Occam's razor.
Maximum parsimony, enumerates differences between sequences and explicitly finds the tree that can be explained by the smallest number of changes, where each difference is a single change and no difference is no change. Distance-based methods estimate expected number of changes between each pair of sequences treating the process of changes as some sort of radioactive decay, i.e. the absence of difference does not mean the absence of change (changes can be followed by a back-substitution to the original state), and a difference may represent multiple changes at this position. Algebraic transformations of these distances result in a tree with certain topology. Maximum likelihood methods attempt to explicitly model substitution process and find a tree that gives the highest probability to the observed data. Bayesian methods, while quite related to maximum likelihood and use similar models to describe substitution process, do the opposite and find the tree, the observed data give the highest probability to. In any case, these 4 types of methods represent all that is available for us to use. We've chosen one software implementation per method: TNT for maximum parsimony, BioNJ for distance, PhyML for maximum likelihood and MrBayes for Bayesian methods. These methods are freely available either for download (TNT, MrBayes) or as servers.
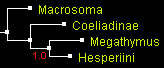
The only tree that was an output of all programs for all genes was the tree 1 that positions Coeliadinae at the base (rooted with Macrosoma). First, we show the trees from combined data of all 4 genes (two nuclear and two mitochondrial), reconstructed by 4 different methods. All the trees look identical with close to 100% statistical support:
| Trees built from 4 genes combined by 4 methods (displayed with ATV) | |||
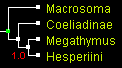 |
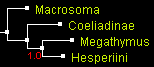 |
 |
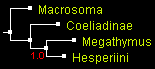 |
| parsimony: TNT | distance: BioNJ | likelihood: PhyML | Bayesian: MrBayes |
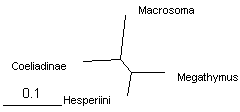
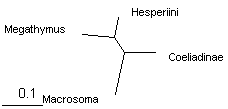
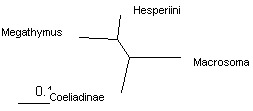
These trees can be shown as unrooted to emphasize relative branch lengths. We see that the longest branch is Macrosoma, as expected for an outgroup. The shortest branch is Hesperia, and the other two branches are about equal length. Thus long branch attraction artifacts are unlikely to occur. Notably, all 4 methods give the same tree.
| The same trees from 4 genes combined built by 3 methods (displayed with TreeView) | |||
 |
 |
 |
|
| distance: BioNJ | likelihood: PhyML | Bayesian: MrBayes | |
All 4 genes were also analyzed individually by 4 programs. All the trees were again identical, most tree were with strong (>90%) statistical support. The lowest supported tree (~60%) was a parsimony tree (TNT) for the NADH dehydrogenase subunit 1 (ND1) gene, which is the shortest gene in the sample and thus is not expected to be statistically significant:
| Trees built from Elongation factor-1 α (EF1) gene by 4 methods (displayed with ATV) | |||
 |
 |
 |
 |
| parsimony: TNT | distance: BioNJ | likelihood: PhyML | Bayesian: MrBayes |
| Trees built from wingless (wg) gene by 4 methods (displayed with ATV) | |||
 |
 |
 |
 |
| parsimony: TNT | distance: BioNJ | likelihood: PhyML | Bayesian: MrBayes |
| Trees built from cytochrome c oxidase subunit I (COI) gene by 4 methods (displayed with ATV) | |||
 |
 |
 |
 |
| parsimony: TNT | distance: BioNJ | likelihood: PhyML | Bayesian: MrBayes |
| Trees built from NADH dehydrogenase subunit 1 (ND1) gene by 4 methods (displayed with ATV) | |||
 |
 |
 |
 |
| parsimony: TNT | distance: BioNJ | likelihood: PhyML | Bayesian: MrBayes |
In addition to the analysis by software, we illustrate positions in the alignment of each gene (and combined genes) that explain why this tree was obtained.
Available aligned EF1-a partial DNA sequences comprise 749 nucleotides at positions present in all 4 species. Multiple alignment is shown below. In most positions, the same nucleotide is present in each of the 4 sequences. E.g. "A" in position 1, "G" in position 2, "A" in position 6 etc. These invariant positions are marked by asterisks below the alignment, positions with at least one difference are highlighted in light-gray:
CLUSTAL W (1.81) multiple sequence alignment 1.Macrosoma_tipulata AGAAGATTGGTTACAACCCAGCTGCCGTCGCTTTCGTACCCATTTCTGGCTGGCACGGAGACAACATGCTGGAGCCCTCTACCAAAATGCCTTGGTTCAAGGGATGGCAAGTTGAGCGCAAGGAGGGCAAGGCTGAAGGCAAGTGCCTCATTGAGGCTCTGGATGCCATCCTGCCCCCTGCTCGTCCCACAGACAAGGCCCTTCGTCTTCCCCTGCAAGACGTATACAAAATCGGCGGTATCGGAACGGTGCCGGTAGGTAGAGTAGAAACTGGTATCCTGAAACCCGGTACCATCGTCGTTTTCGCCCCTGCCAACATTACCACTGAAGTCAAGTCCGTTGAGATGCACCACGAAGCCCTCCAAGAGGCCGTACCCGGAGACAATGTTGGTTTCAACGTAAAGAACGTGTCTGTCAAGGAGTTGCGTCGTGGATACGTTGCCGGTGATTCCAAGAATAACCCACCTAAGGGTGCATCCGACTTCACTGCACAGGTCATCGTGCTTAACCACCCTGGTCAAATTTCCAATGGTTATACACCCGTGTTGGATTGCCACACTGCCCACATAGCCTGCAAATTCGCTGAAATCAAGGAGAAGGTCGACCGTCGTACTGGTAAATCCACTGAGGAGAACCCCAAATCTATCAAATCTGGTGACGCTGCCATTGTCAACTTGGTTCCCTCCAAGCCCCTGTGTGTGGAGTCCTTCCAGGAATTCCCTCCCCTTGGTCGTTTCGCC 2.Coeliades_forestan AGAAGATCGGTTACAACCCCGCTGCTGTCGCTTTCGTTCCCATTTCTGGCTGGCACGGAGACAACATGCTGGAGCCATCCACCAAAATGCCCTGGTTCAAGGGATGGTTGGTTGAGCGCAAGGAAGGAAAGGCCGAAGGTAAATGCCTCATTGAAGCTTTGGACGCCATCCTGCCGCCTGCTCGCCCCACAGACAAGGCCCTCCGTCTTCCCCTGCAGGACGTCTACAAAATCGGTGGTATTGGTACGGAGCCCGTAGGCAGAGTTGAAACTGGTGTCCTCAAACCCGGTACCATTGTTGTCTTTGCCCCGGCCAACATCACCACTGAAGTYAAGTCTGTGGAGATGCACCACGAAGCCCTCCAAGAGGCTGTCCCCGGAGACAACGTCGGTTTCAACGTTAAGAACGTCTCCGTCAAGGAATTGCGTCGTGGTTACGTCGCTGGTGACTCCAAGAACAACCCACCCAAGGGTGCCGCAGACTTCACAGCKCAGGTCATCGTGCTTAACCACCCCGGACAAATCTCCAACGGATACACACCAGTGTTGGATTGCCACACCGCTCACATTGCCTGCAAGTTCGCAGAGATCAAAGAGAAGGTTGACCGTCGTACTGGTAAATCTACTGAAGACAATCCCAAATCCATCAAGTCTGGTGATGCCGCCATTGTCAACCTCGTGCCCTCCAAGCCTCTGTGCGTGGAGTCCTTCCAGGAATTCCCACCCCTCGGTCGTTTCGCC 3.Megathymus_streckeri AGAAAATAGGTTATAACCCAGCTGCCGTCGCTTTCGTACCCATTTCTGGCTGGCACGGAGATAACATGCTGGAGCCATCTACCAAGATGCCTTGGTTCAAGGGATGGAATGTTGAGCGTAAGGAAGGTAAGGCTGAAGGGAAATGCCTCATTGAGGCTCTTGATGCCATCCTACCCCCAGCTCGTCCCACAGACAAAGCCCTCCGTCTTCCTCTGCAGGATGTCTACAAAATCGGTGGTATTGGTACAGTGCCAGTAGGCAGAGTTGAGACTGGAATCCTGAAGCCTGGTACCATTGTTGTATTTGCTCCCGCTAATATCACTACTGAAGTAAAATCTGTGGAAATGCACCACGAGGCTCTCCAGGAGGCTGTGCCTGGAGACAATGTAGGATTCAACGTAAAGAATGTATCTGTTAAAGAATTGCGTCGTGGATACGTCGCCGGTGATTCCAAAAACAACCCACCAAAGGGTGCCGCAGATTTCACAGCACAAGTCATTGTGCTCAACCATCCTGGTCAAATTTCCAACGGTTATACACCAGTATTGGATTGCCACACTGCTCACATTGCCTGCAAATTTGCAGAGATTAAAGAGAAGGTTGACCGTCGTACTGGTAAATCTACTGAGGAGAATCCTAAATCTATCAAGTCTGGTGATGCGGCCATTGTTAACTTAGTACCTTCCAAGCCTCTGTGTGTAGAATCCTTCCAGGAGTTTCCACCCCTCGGTCGTTTCGCT 4.Hesperia_leonardus AGAAGATTGGTTACAACCCAGCTGCCGTCGCTTTCGTACCCATTTCTGGCTGGCACGGAGACAACATGTTGGAGCCATCTACCAAGATGCCTTGGTTCAAGGGATGGAATGTTGAGCGCAAGGAAGGTAAGGCTGAAGGCAAATGCCTCATTGAGGCCCTGGACGCCATCCTGCCCCCAGCTCGTCCCACAGACAAAGCCCTCCGTCTTCCCCTGCAGGACGTCTACAAAATCGGTGGTATTGGTACAGTGCCCGTAGGCCGAGTGGAAACTGGAATCCTGAAGCCTGGTACCATTGTTGTGTTCGCCCCCGCCAACATCACCACTGAAGTCAAATCTGTGGAGATGCACCACGAGGCTCTCCAAGAGGCTGTGCCTGGAGACAATGTAGGTTTCAACGTAAAGAACGTCTCTGTCAAGGAATTGCGTCGTGGCTACGTCGCCGGTGACTCCAAGAACAACCCACCCAAGGGTGCCGCAGACTTCACTGCACAAGTCATTGTGCTCAACCACCCTGGTCAAATTTCCAACGGTTACACGCCTGTGTTGGATTGCCACACTGCCCACATTGCCTGCAAATTCGCAGAAATCAAAGAGAAGGTGGACCGTCGTACTGGTAAATCGACTGAGGAAAATCCCAAATCTATCAAATCTGGTGATGCCGCTATTGTAAACTTAGTACCTTCCAAGCCTCTGTGTGTAGAATCCTTCCAGGAGTTCCCACCTCTCGGACGTTTCGCC **** ** ***** ***** ***** *********** *********************** ****** ******* ** ***** ***** *************** ******** ***** ** ***** ***** ** *********** ** * ** ******** ** ** ***** *********** ***** ******** ***** ** ** *********** ***** ** ** * *** ***** **** ** ***** **** ** ** ******** ** ** ** ** ** ** ** ** ** ******** ** ** ** ** *********** ** ***** ***** ** ** ******** ** ** ******** ***** ** ** ** ** ** *********** ***** ** ***** ***** ** ******** ******** * ** ***** ** ** ***** ***** ***** ** ** ***** ***** ** ** ** ** ** ************** ** ***** ******** ** ** ** ** ** ******** ******************** ***** ** ** ** ***** ***** ******** ** ** ***** *** * ** ** ******** ***** ** ** *********** ** ** ** ** ** ********
Since positions with identical nucleotides do not directly tell us which sequence is a neighbor of which (there are no changes in such positions), these positions are not considered in parsimony analysis and can be removed from the alignment. As a result, we get the following alignment of 144 positions:
1.Macrosoma_tipulata 31 GTCACACCCTATCAACGCTCGGTCGTGCTTGTCACACCAGTGTAAATAGACCCTCCTCCTCCGCTGACACACTTTACGTCGGATCTGTTATCCTAGCTCTTTTTTACGTCAACTACGCCGGCCTACTCCTGTCCTGGACTCTTC 31 Macrosoma_tipulata 2.Coeliades_forestan 29 GCCCTTCCACACTTGCAACTAATTGCGGTCGCCGCCTTTGACCATATGCACTTCTCGCCCCYGTGGACATCCCCTTCCCCGATCTCGCCCGACAKGCTCCACCACAAGCTTGCAGCATTACTCCGTCCCCCGCTCGGACACCTC 29 Coeliades_forestan 3.Megathymus_streckeri 28 AATACATCATGTAATTATTGAGTCTTACATACTGTCTTTATACATGAAGGTTTATTCTTCTAATGAGTGTGTTAAATATTAAACCTACACGATAAATCTTTTCTTAAATTTATAGTATTGGTTTGTGCTTAATTTAAGTACCTT 28 Megathymus_streckeri 4.Hesperia_leonardus 7 GTCACACTATGTAATCATTCAGCCGCGCATACCGCCTTTATCCCGAAAGGTTTGCCCCCCCCATGGGTATGTTATACCTCGACCCCGCCCGACTAATCCTTTCTCGTGTCTACAACAGGGATCTATCTATAATTTAAGCATCAC 7 Hesperia_leonardus
Nucleotides unique to each sequence (i.e. position has the same nucleotide in 3 sequences, but it is different in the 4th sequence, =semi-invariant position) are highlighted in gray and their number is shown before (and after) each sequence on yellow background. In the simplest incarnation of the maximum parsimony method we need to find the tree that can be explained by the smallest number of nucleotide substitutions. These positions with nucleotides unique to one sequence can be explained by a single substitution: from the nucleotide present in remaining 3 sequences to a nucleotide present in the 4th sequence. E.g. the 1st position {G,G,A,G} can be explained by a single substitution of G to A. Since A is present only in one sequence, this substitution should be assigned to (=presumable happened on) the tree edge (=branch) leading the sequence with A as a leaf, i.e. the G → A substitution happened on the terminal branch leading to Megathymus streckeri. Because for all 3 tree tress positions with nucleotides highlighted in gray can be explained with a single substitution, these terminal branches do not directly assist us in topology selection. Removing semi-invariant positions from the alignment, we get even shorter alignment of only 49 positions:
1.Macrosoma_tipulata TACACCTTGGGATACTCTCGACACTGATTTGCTTCCACCGATCGTCGGA 3 Macrosoma_tipulata 2.Coeliades_forestan CATGATCTGGCTTACCTGYGACCCCCTCCAGCTCATGTTCGCCCGCGGA 5 Coeliades_forestan 3.Megathymus_streckeri AGATTGTAAAATAGTATCAAGTGTAAATAAATCTATGTTGGGTAATAAG Megathymus_streckeri 4.Hesperia_leonardus TGATTCCAAACGAGTGCCCAGTGTACCCCTATCCTCAGGAACAAATAAG 18 Hesperia_leonardus
Next, we highlight "2+2" positions, i.e. positions with the same nucleotide in 2 sequences and a different nucleotide common to the other two sequences. There are 3 types of such positions: type 1 (highlighted green) has the same nucleotide in sequences 3 and 4, type 2 (highlighted red) has the same nucleotide in sequences 1 and 3, and type 3 (highlighted blue) has the same nucleotide in sequences 2 and 3. Number of these positions is shown highlighted yellow to the right of the sequences: type 1 after sequence 4, type 2 after sequence 1 and type 3 after sequence 2. These positions can also be explained by a single substitution, but on the middle tree branch, and only in case the middle branch separates the species pairs with the same nucleotide in a pair, but different nucleotide between the pairs. I.e. the second position in the alignment {A,A,G,G} can be explained by a single change (either A → G, or G → A) in case the tree 1 (Megathymus is joined with Hesperia) is correct. For the other two tree topologies (Megathymus joined with Macrosoma, or Megathymus joined with Coeliades), minimum 2 changes are necessary. E.g. for the tree 2 (Megathymus is joined with Macrosoma) we need a change A → G in Megathymus and another change A → G in Hesperia, which makes it 2 changes. Alternatively, this position can be explained by a change G → A in Macrosoma and another change G → A in Coeliades – also 2 changes. Thus the position {A,A,G,G} votes for the grouping of Megathymus with Hesperia, as the smallest number of changes are needed to explain it for the tree 1. Positions of type 1 (green) support the tree 1 (Megathymus joined with Hesperia), positions of type 2 (red) support the tree 2 (Megathymus placed at the base), and positions of type 3 (blue) support the tree 3 (Hesperia placed at the base). These "2+2" positions are called parsimony-informative, as the smallest number of changes assigned to them depends on the tree topology. The position type with the highest vote wins parsimony. Apparently, in this case 18 > 5 > 1, thus green positions (type 1) win, and the most parsimonious tree is 1, in which Coeliades is the basal group. It has to be mentioned that the smallest number of changes assigned to positions without highlight in the alignment (e.g. {T,C,A,T}) does not depend on tree topology (for {T,C,A,T} it is always 2), and can be ignored in this method.
We see that the tree 1 is supported by an excess of 18 − 5 = 13 positions, which is about 3 times the number of positions supporting alternative grouping (Tree 3:), and finally the tree 2 (Megathymus basal) is the least parsimonious option.
The data for other 3 genes are presented similarly: full alignment with variable positions highlighted, alignment with removed invariant positions with semi-invariant positions highlighted, and the alignment with invariant and semi-invariant positions removed, positions supporting different tree topologies are highlighted in colors assigned to these topologies:
CLUSTAL W (1.81) multiple sequence alignment Macrosoma_sp. GTGCACGGTCAAGACCTGTTGGATGCGGCTGCCGAGCTTGCGCTCTGTGGGCGACGCGTTGAAGGAGCGCTTCGACGGCGCCTCGCGGGTGATGATGCCTAACACGGACCTGGAGGCGCCGTCCCAACGGAACGACGCGGCGCCGCACAGAGTGCCCCGGCGGGACCGCTACCGGTTCCAACTGCGGCCGCACAACCCCGAGCACAAGTCGCCAGGCGCGAAGGACCTGGTCTACTTGGAGTCGTCGCCGGGCTTCTGCGAGAAGAACCCCAGGCTCGGCATTCCCGGCACCCACGGGCGCGCATGCAACGACACGAGCATCGGCGTCGACGGCTGCGACCTAATGTGCTGCGGCCGCGGTTACAGAACTGAGACTATGTTCGTTGTGGAAAGATGCAAC Coeliades_forestan GTGCACGGTGAAGACGTGCTGGATGAGGCTGCCGAGTTTCCGCTCCGTGGGCGACGCGTTGAAGGACCGCTTCGACGGGGCGTCACGGGTCATGATGCCCAACACCGACCTGGAGGCGCCCGTGCAGCGGAACGACGCCGCGCCGCACCGAGTCCCGCGAMGGGATCGCTACAGATTCCAACTCCGGCCTCACAATCCGGACCACAAGTCTCCGGGGGCCAARGACCTCGTGTACCTGGAATCCTCGCCGGGTTTCTGCGAAAAGAACCCGAGACTCGGCATTCCCGGGACGCACGGGCGCGCCTGCAACGACACGAGCATCGGCGTCGACGGCTGCGACCTCATGTGTTGCGGCCGAGGCTACCGGACSGAGACCATGTTCGTGGTGGAACGGTGCAAC Megathymus_streckeri GTGCACGGTGAAGACCTGCTGGATGAGGCTACCGAGTTTCCGTTCGGTGGGGGACTCGCTGAAGGATCGCTTCGACGGGGCCTCGCGWGTTATGATGCCCAACACGGACCTGGAAGCACCCGTGCAACGGAACGATGCGGCTCCTCACAGGATACCCCGGAGAGATCGATATCGTTTCCAACTCCGACCGCACAACCCAGATCACAAATCTCCGGGGATCAAAGACCTCGTGTATCTGGAACCGTCGCCGGGCTTCTGCGAAAAGAACCCGCGGCTGGGCATTCCCGGCACACATGGGCGGGCGTGCAACGACACGAGTATCGGTGTCGACGGCTGCGACCTCATGTGTTGCGGTCGCGGCTACAGGACCGACACTATGATTGTCGTGGAACGGTGTAAT Hesperia_leonardus GTGCACGGTGAAGACCTGCTGGATGAGACTGCCGAGTTTCCGTTCTGTGGGCGATGCGTTGAAGGATCGCTTCGACGGGGCCTCTCGAGTGATGATGCCCAACACGGACCTGGAAGCGCCCGTACAACGGAACGATGCCGCTCCGCAYCGGGTACCCCGGCGWGATCGGTATCGTTTCCAGCTCCGACCGCACAACCCGGACCACAAATCTCCGGGYGCCAAGGACCTCGTGTATTTAGAACCGTCACCGGGTTTCTGCGAAAAGAACCCTCGGCTCGGCATTCCCGGCACACACGGGCGGGCCTGCAACGACACGAGCATCGGCGTCGACGGCTGCGACCTCATGTGTTGCGGCCGCGGCTACAGAACCGAGACTATGTTTGTTGTAGAACGATGTAAT ********* ***** ** ****** * ** ***** ** ** ** ***** ** ** ******* *********** ** ** ** ** ******** ***** ******** ** ** ** ******** ** ** ** ** * * ** ** * ** ** ** * ***** ** ** ** ***** ** ** ***** ** ** ** ** ***** ** ** * ** * ** ***** ******** ******** * ** *********** ** ** ***** ** ************** ***** ***************** ***** ***** ** ** *** * ** ** ** *** * ** ** *** * ** **
Macrosoma_sp. 23 CCTCGGCGCTCCGTGCCGGGTGGGGTCCACGGGCAAGGCGCGCCCCGAGGGCCGGGACGCGGGCCTGGTGGCGCAGCCCCCACCACCCTAATGTTCTGAACC 23 Macrosoma_sp. Coeliades_forestan 15 GGCAGGTCCCCCGTCGGAGCCCGGCGTGGCCGGCCAGCGAMGTCCAAACGTTGCGTGGGCCRCGCCGATCGTAGAACGGCCCCCCTCACCGSGCTCGGCGCC 15 Coeliades_forestan Megathymus_streckeri 16 GCCAGATCTGGCTCTGCGWTCGAACGTGATGTTCAGAACGAATATCTACAGCATATGGATCACGTCGACGGCAGCGGCATGGTTCTTCCAGCCTATCGCGTT 16 Megathymus_streckeri Hesperia_leonardus 7 GCCAAGTCTTCTGTTGCTAGCGAGCGTAATCTGYCGGACGCWTGTCTGCAGCGCATGYGCCGCGTTAACGATATCGCCACGCCCCTCCCAACGTTTTACATT 7 Hesperia_leonardus
Macrosoma_sp. CTGGGGGCCGGAAGCGCCGGCGGCGCTTCCACCAATCTACC 3 Macrosoma_sp. Coeliades_forestan CCCAGCGGCCGCACMGCCAGGCGGRCCTTGAGCCGSCGGCC 3 Coeliades_forestan Megathymus_streckeri TGTGWTAGTGTAGAAAATTAATAGATCCCGCAGGGCTCGTT Megathymus_streckeri Hesperia_leonardus TTTTAGAATCTCGACWGTTAGCAYGTTCTTCAGCACTTATT 15 Hesperia_leonardus
The strongest support (15 positions) is for the tree 1.
CLUSTAL W (1.81) multiple sequence alignment Macrosoma_sp. ACTTGGATTTATTGTTTGAGCTCATCATATATTTACTGTAGGTATAGATATTGATACTCGAGCTTATTTCACATCAGCAACTATAATTATTGCTGTTCCTACTGGAATTAAAATCTTTAGTTGACTAGCTACTCTTCATGGAACTCAAATTAATTATAGTCCTTCTATATTATGAAGATTAGGATTTGTATTTTTATTTACAGTAGGGGGATTGACAGGAGTAATTTTAGCTAATTCTTCTATTGATATTACTTTACATGATACTTATTATGTAGTAGCTCATTTTCATTATGTTTTATCTATAGGAGCTGTATTTGCCATTATAGGAGGATTTATTCATTGATACCCTTTATTTACAGGATTAATAATAAACCCTTATCTATTAAAAATTCAATTTTTTTCTATATTTATTGGAGTAAATTTAACTTTTTTTCCTCAACA-TTTTTTAGGTCTAGCAGGTATACCTCGTCGATATTCAGATTATCCTGATAGTTTTATTTCTTGAAATATTATTTCTTCATTTGGTTCTTATATTTCTTTATTATCAACAATATTAATAATAATTATTATTTGAGAATCGATGATTAATCAACGAATTATTTTATTTTCAATAAATATACCTTCCTCAATTGAATGATTCCAAAATCTTCCTCCTGCAGAACATTCATATAATGAATTACCTATTTTAAGAAATTTCTAATATGGCAGATTATATGTAATGGATTTAAACCCCATTTATAAAGGATTATCCTTTTTTTAGAA-ATGGCAACTTGATCAAATTTTAATTTACAAAATGCAGCATCCCCTTTAATAGAACAAATTATTTTTTTTCATGATCATACATTAGTTATTTTATTAATAATTACAATTTTAGTAAGTTATTTAATAG Coeliades_forestan ATTAGGATTTATTGTTTGAGCTCATCATATATTTACTGTCGGAATAGATATTGATACTCGAGCATATTTTACTTCAGCCACAATAATTATTGCGGTACCAACTGGAATTAAAATTTTTAGTTGATTAGCTACTCTTCATGGTACTCAAATTAATTATAGTCCTTCTATACTTTGAAGATTAGGATTTGTATTCTTATTTACAGTAGGTGGATTAACAGGAGTAATTTTAGCCAATTCTTCTATTGATATTATACTTCATGATACTTACTATGTAGTAGCTCATTTCCATTATGTTCTATCTATAGGAGCAGTATTTGCTATTTTAGGGGGATTTATTCACTGATATCCTTTATTTACAGGATTATCTCTTAACCCTTATTTATTAAAAATTCAATTTATCTCTATATTTTTAGGAGTAAATTTAAC-TTTTTTCCTCAACACTTTTTTAGGATTAGCTGGTATACCTCGTCGTTATTCAGATTATCCTGACAGTTATATTTCATGAAATATTATTTCTTCTTTAGGTTCTTATATTTCTTTATTATCAATAATAATAATAATAATTATTATTTGAGAATCTATAATTAATCAACGAATTGTTTTATTTTCTCTTAATATACCTTCTTCTATTGAATGATATCAAAATTTACCTCCTGCTGAACACTCATATAATGAATTACCTATTTTAAGAAACTTCTAATATGGCAGATTATATGTAATGGATTTAAACCCCATTTATAAAGGAATATCCTTTTTTTAGAA-ATGGCAACATGATCAAATTTTAATTTACAAAATAGAGCTTCTCCATTAATAGAACAAATTATCTTTTTTCATGATCATACTTTAGTTATTTTAATTATAATTACTATTTTAGTAGGATATTTAATAA Megathymus_streckeri ATTAGGATTTATTGTTTGAGCTCATCATATATTTACTGTAGGTATAGATATTGATACACGAGCTTATTTTACTTCAGCAACTATAATTATTGCAGTACCAACTGGTATTAAAATTTTTAGTTGATTAGCTACACTTCATGGAACAAAAATTAATTATAGACCTTCTATATTATGAAGATTAGGATTTGTATTTTTATTTACTGTAGGAGGATTAACAGGAGTAATTTTAGCTAATTCTTCTATTGATATTACTCTTCATGATTCCTATTATGTAGTAGCTCATTTTCATTATGTATTATCTATAGGAGCAGTATTCGCTATTTTCGGAGGATTTATTCATTGATATTCATTATTTACCGGATTAACTTTAAATTCTTATTTATTAAAAATTCAATTTATTTCAATATTTTTAGGAGTAAATTTAACTTTTTTTCCTCAACA-TTTTTTAGGTTTATCAGGAATACCTCGACGATATTCTGATTACCCTGATAGTTATTTATCTTGAAATATTATCTCATCATTTGGATCTTATATTTCTTTATTTTCAATAATAATAATAATAATAATTATTTGAGAATCATTTATTAATCAACGTTTAATTTTATTTTCATTAAATATATCTTCTGCTATTGAATGACTTCAAAATTTCCCCCCTAATGAACATTCTTATAATGAATTACCTATTTTAAGAAATTTCTAATATGGCAGATTATATGTAATGGATTTAAACCCCATTTATAAAGGATTATCCTTTTTTTAGAATATGGCAACATGATCTAATTTTAATTTACAAAATAGAGCTTCACCTTTAATAGAACAAATTATTTTCTTCCATGATCATACCTTAATTATTTTAATTATAATTACCATCTTAGTTAGATATATAATAA Hesperia_leonardus ATTAGGATTTATTGTTTGAGCTCATCATATATTTACAGTTGGTATAGATATTGATACTCGAGCTTATTTTACTTCAGCTACTATAATTATTGCAGTACCTACAGGAATTAAAATTTTTAGTTGATTAGCAACTCTTCATGGAACTCAAATTAATTATAGCCCTTCTATATTATGAAGATTAGGATTTGTATTTTTATTTACAGTAGGAGGATTAACTGGAGTAATTTTAGCTAATTCTTCAATTGATATTACTCTTCATGATTCTTATTATGTAGTTGCTCATTTTCATTATGTTTTATCAATAGGAGCTGTATTTGCTATTTTTGCAGGATTTATTCATTGATACCCATTATTTACAGGATTACCAATAAATAATTATATATTAAAAATTCAATTTATTTCAATATTTATTGGAGTTAATTTAACATTTTTCCCACAACA-TTTTTTAGGTTTAGCTGGAATACCACGACGTTATTCTGACTATCCTGATAGTTATCTTTCATGAAATGTAATTTCATCTTTTGGATCTTATATTTCTTTATTATCTATAATAATAATAATAATAATTATTTGAAAATCAATAGTTTATCAAAATATTGTATTATTTTCTTTAAATATATCATCTTCTATTGAATGACTTCAAAATTTACCCCCAACTGAACATTCATATAATGAATTACCTATTTTAAGAAATTTCTAATATGGCAGATTATATGTAATGGATTTAAACCCCATTTATAAAGGTTTATCCTTTTTTTAGAATATGGCAACATGATCTAATTTAAATCTTCAAAATAGAGCTTCTCCTTTAATAGAACAAATTATTTTTTTCCATGATCATACTTTAATTATTTTAATTATAATTACTATTTTAGTTAGTTATATAATAA * * ******************************** ** ** ************** ***** ***** ** ***** ** *********** ** ** ** ** ******** ********* **** ** ******** ** ************* ********* * ******************** ******** ***** ***** ** ************** ******** ********** * ****** * ** ******** ******** ******** **** ******** ***** ** *** * * *********** ***** * ******** ****** * ** **** ***************** * ** ****** * ***** ******** ***** ** ***** ********* ** * ** ***** ** ** ***** ** ** ***** **** * * ** ****** * ** ** ** ** ** ***************** ** * **** ********** ********* **** * ** ***** * * ******** * ****** * ** * ********* ****** * ** ** ***** ** ************************** ************************************************** **************** ******** ***** ***** *** * ****** *** ** ** ***************** ** ** *********** *** ******** * ******** ** ***** * *** *****
Macrosoma_sp. 30 CTTATTTCAATTTTTACCTTATCTTATAGGATTCTTAATTATTTTTTCAAGATCCTAATAAACCCCTTTATATTT-TCGATTTAATTTTATTATTTATTAACTTGGAGAACGAATATAAACTCTATTCCTTTGCATATAT-TATTAGCACTTTTAGTAATAATTG 30 Macrosoma_sp. Coeliades_forestan 29 TATCATATTCAGAATATTTTTTCTCTCATAACTTACTATCACTCTATTTAGGCTCTATCTCTCCCTACTTAA-TTCATGTTTTTATTCAATAATTTTATAATATGTAAAACGAATGTTCTCTTTTTATTATTGCTCACAA-AATTAAGTTACTTTGATTTAGATA 29 Coeliades_forestan Megathymus_streckeri 24 TATATATTTATAAATTTTTAAAAATATTAAATTCTCTTCTATATTACTTCGATTTACACTTATTCTATATAATTT-TTTAATAATTCTATATATCAATATATAAGATTAACGTTAATATATTTGTCTTTCCTAATTTTATTATTTAAGTATTCCCAATCCTAAAA 24 Megathymus_streckeri Hesperia_leonardus 29 TAATTTTTTTTAATAATTATATCCTATAAATTACTCTTTTTTTTATTTTTCATCCAACCAAATAAAATAATTACA-TTGTAAATTCTTACTAGATATTAATTAAAAAAGTAATATGATTATATTTCTTTACAACTTATTTTATACTAGTTTTTCTAATTTTATAA 29 Hesperia_leonardus
Macrosoma_sp. AATTTGATACTAAACCCTATTATTAAATTATTGGAAAACTTTG-ACTAGAATT 6 Macrosoma_sp. Coeliades_forestan CCGATTAAATTTTCCCTTTA-TTTTAAATTTTTAAGTCCTATG-ATTTGTAAT 7 Coeliades_forestan Megathymus_streckeri AAAAAATACTAATTTTTATATAAAATTTAAAAATTAATTCCCATTACCACTAA Megathymus_streckeri Hesperia_leonardus TTATCATTTCACAATAAAATATAATTCAATAAAATGTTTCACATTTCTATTTA 21 Hesperia_leonardus
21 positions for the tree 1 vs. three times less for other trees.
CLUSTAL W (1.81) multiple sequence alignment Macrosoma_semiermis AGTAATTGGGATTTTACAGCCTTTTTCTGATGCGATTAAATTATTTAGTAAAGAAATAATTTATCTTAATTTATCTAATTATATGATTTATTGTTTTTCTCCTATCTTTAGATTTATTTTATCTTTAATAATGTGAATAATAATTCCTTATTATTTTAATATATTTWAGTTTARATTAGG Choaspes_benjaminii GTATATAGGTTTATTACAGCCTTTCTCAAATGCTATTAAATTATTTACTAAGGAGCAAATATATTTAAATTATTCTAATTATATATCAAATTATTTTTCTCCAATTATTAGTTTTATTTTGTCTTTAATGATTTGAATATTAATTCCTTATTATTTTAATATAGTTAGTTTTAATTTAGG Megathymus_yuccae TTTTATAGGTTTATTACAGCCTTTTTCTGATGCTATTAAATTATTTAGAAAGGAGCAAGTTTATTTATTAAATGCTAATTATTTAGTTTATTATTTTTGTCCTATTATTGGTTTTATTTTWTCTTTATTGATTTGAGTATTAGTTCCTTATTATTTTAATTTAATTAGATTTAATWNAGG Hylephila_phyleus ATTTATGGGTTTATTACAACCTTTTTCTGATGCTATTAAATTATTTACAAAAGAACAAACTTATTTATNGAATTCTAATTATTTAATTTATTATTTTTGTCCTATTGTTAGATTTATTTTATCATTAATAATTTGAGTTTTAATACCWTATTATTTTAATTTAATTAGATTTAATTTGGG ** ** * ***** ***** ** **** ************* ** ** * *** * ******** * *** ***** *** ** ** * ******** ** *** * ** *** * ** * ** ************ ** ** **** **
Macrosoma_semiermis 21 AGTATGATGTTGGGTAAATATTCTAATTTATAGATTTGCTCTAAATAAGAAAATTATWAGRATTA 21 Macrosoma_semiermis Choaspes_benjaminii 9 GTATATTAGCAATCTGGCAATATAAATTATTAATCAAACATAATGTAGTAATATTAGAGTATTTA 9 Choaspes_benjaminii Megathymus_yuccae 7 TTTTATTAGTTGTGAGGCAGTTTATTAAATGTAGTTTAGTTAGTWTTGTGATGTTTAAGAATWNA 7 Megathymus_yuccae Hylephila_phyleus 7 ATTTGTTAATTGTCAAACAACTTATNGAATTTAATTTAGTTGAAAAAATGTTAAWTAAGAATTTG 7 Hylephila_phyleus
Macrosoma_semiermis ATGTAAAATTAACTAAAAATG 1 Macrosoma_semiermis Choaspes_benjaminii GACTGGAATTATCATGGAAGT 4 Choaspes_benjaminii Megathymus_yuccae TAGAGGTTAATGGATWGGTAA Megathymus_yuccae Hylephila_phyleus AGCAAATNGATAGGAAAGTAA 7 Hylephila_phyleus
Only 7 positions for the tree 1, vs. 4 or 1 position. This is the shortest gene segment and statistical support for the tree is the lowest.
Finally, the combined over 4 genes segment with parsimony-informative positions is shown:
Macrosoma 13 AATTTGATACTAAACCCTATTATTAAATTATTGGAAAACTTTG-ACTAGAATTTACACCTTGGGATACTCTCGACACTGATTTGCTTCCACCGATCGTCGGACTGGGGGCCGGAAGCGCCGGCGGCGCTTCCACCAATCTACCATGTAAAATTAACTAAAAATG 13 Macrosoma Coeliadinae 19 CCGATTAAATTTTCCCTTTA-TTTTAAATTTTTAAGTCCTATG-ATTTGTAATCATGATCTGGCTTACCTGYGACCCCCTCCAGCTCATGTTCGCCCGCGGACCCAGCGGCCGCACMGCCAGGCGGRCCTTGAGCCGSCGGCCGACTGGAATTATCATGGAAGT 19 Coeliadinae Megathymus AAAAAATACTAATTTTTATATAAAATTTAAAAATTAATTCCCATTACCACTAAAGATTGTAAAATAGTATCAAGTGTAAATAAATCTATGTTGGGTAATAAGTGTGWTAGTGTAGAAAATTAATAGATCCCGCAGGGCTCGTTTAGAGGTTAATGGATWGGTAA Megathymus Hesperiini 61 TTATCATTTCACAATAAAATATAATTCAATAAAATGTTTCACATTTCTATTTATGATTCCAAACGAGTGCCCAGTGTACCCCTATCCTCAGGAACAAATAAGTTTTAGAATCTCGACWGTTAGCAYGTTCTTCAGCACTTATTAGCAAATNGATAGGAAAGTAA 61 Hesperiini
61 positions (highlighted green) support the tree 1, and over three times fewer positions support other trees.
Conclusions:
| Male genitalia of the species considered here and their close relatives. |
 |
With the help from Brian Banker, Stan Gorodenski, Dave Ferguson, Jon Pelham and others, we assembled the following list of 80 characters.
Character is a question that can be answered either "yes" or "no". Character states are: "+" - answer "yes", "-" - answer "no", "?" - answer "don't know". Taxa used: Mac - Macrosoma tripulata, Coe - Coeliades forestan, Meg - Megathymus streckeri, Hes - Hesperia leonardus. Character matrix: # character Mac Coe Meg Hes 1. Clubbed antennae - + + + 2. Antennae thinly narrow towards the tip + + - - 3. Antennae with apiculus - + - + 4. Clubbed antennae without apiculus - - + - 5. Antennae shorter than 1/2 costa - - + + 6. palpi are longer foreleg tibiae - + - + 7. 3rd palp segment long, projects frwrd - + - - 8. "Eyelash" present - - + + 9. Two pairs of chaetostomata - - + + 10. Undersized head in relation to body - - + - 11. Nocturnal superposition eyes + - - - 12. Male foreleg not used for walking + - - - 13. Sexes alike + + - - 14. Wings coupled (frenulum/retinaculum) + - - - 15. FW M2 vein joins closer to M3 - - + + 16. FW cell longer than base-to-tornus dis - + - - 17. Male with FW stigma - - - + 18. apex-tornus-base FW angle > 100Deg + + - - 19. HW longer than the abdomen - + - - 20. HW base with a small tuft of hairs - - + + 21. Prominent androconial HW tufts - - + - 22. FW with large postmedian light spots - - + + 23. FW without light spots near the apex + + - - 24. FW discal cell with a spot at the end + - + + 25. HW discal cell lighter than FW d.cell + + - - 26. VHW with a white band - + - - 27. Abdomen curved + - - - 28. Pouched form of the 1st abdom. tergum + - - - 29. Adult body very slender + - - - 30. Tibiae spined - - + + 31. metatibial spurs + + - + 32. All ova fully-formed at emergence - - + - 33. Oviposit mostly on the leaf underside - ? + + 34. Egg upright, Pierid or Nymphalid-like + - - - 35. egg with longitudal "ribs" + + - - 36. Egg with cross-ribs + - - - 37. Horizontally oval ovum shape - - + - 38. Larvae feed on monocots - - + + 39. Genus host plants in several families + + - - 40. larvae feeding on leaves in all stages + + - + 41. Butterfly-like larva + + - + 42. colorless final stage larva - - + - 43. Larvae in cylindrical webbed tunnels - - + + 44. 1st instar with long setae on last sgm - - + + 45. reddish, pigmented first instar - - + - 46. Larva striped, brightly colored - + - - 47. Last intar larva mostly green + - - - 48. Last instar larva mostly brown - - - + 49. larval head black - - + + 50. no anal comb in larvae - - + - 51. Larval head with 2 long scoli + - - - 52. powdering larvae - ? + + 53. larva powdering the cocoon exit - - + - 54. pupae are hold by a "sling" + + - - 55. Butterflylike pupa + - - - 56. Fully articulating chrysalis - - + - 57. Point-style cremaster (vs.flattened) + + - + 58. Pupa with "horn" on thorax + - - - 59. Pupa covered in powder - + + + 60. Stored-fat lifestyle - - + - 61. Boring (in-root) lifestyle - - + - 62. Fast flight - + + + 63. Nectar-feeding adults ? + - + 64. Mating highly reliant on pheromone ? ? + ? 65. Adult basking no jet-pose + + - - 66. Rhopalocera resting posture - + + + 67. Resting posture - wings erect - + - - 68. At times crepuscular adults + + + - 69. Old Word distribution - + - - 70. Known migratory behavior - + - - 71. Uncus long finger-like laterally + + - - 72. Valva rounded at the tip + - - - 73. Gnathos arms almost reach uncus tip - - + + 74. Harpe defined but small - - - + 75. Valva broad, <1/3 of length - - - + 76. Harpe ~1/2 of valva, with many teeth - + - - 77. Valva P-shaped - + + - 78. Penis shorter than valva ? - + - 79. Valva slightly convex ventrally + - - - 80. “Vinculum” W-shaped laterally - + - -
The states for the characters were taken from the following literature and augmented by personal observations of the participants of this discussion.
Warren, A. D., Ogawa, J. R. & Brower, A. V. Z. 2009 Revised classification of the family Hesperiidae (Lepidoptera: Hesperioidea) based on combined molecular and morphological data. Systematic Entomology 34, 467-523.
Scoble, M.J. (1986) The structure and affinities of the Hedyloidea: a new concept of the butterflies. Bull. Brit. Mus. (nat. Hist.) (Ent.), 53: 251-286.
Lourido, G., Silva, N.M., Motta, C.S. 2007. Biological Parameters and Damage by Macrosoma tipulata Hübner (Lepidoptera: Hedylidae), in Cupuaçu tree [Theobroma grandiflorum (Wild ex Spreng Schum)] in Amazonas, Brazil. Neotropical Entomology, 36(1):102-106.
Scoble, M.J., Aiello, A. (1990). Moth-like butterflies (Hedylidae: Lepidoptera): a summary, with comments on the egg. Journal of Natural History, 24(1): 159-164.
Kendall, R.O., (1976). Larval foodplants and life history notes for eight moths from Texas and Mexico. Journal of the Lepidopterists' Society, 30(4): 264-271.
Aiello A. 1992. Nocturnal Butterflies in Panama, Hedylidae (Lepidoptera: Rhopalocera). In: Diomedes Quintero Arias and Annette Aiello (eds.). Insects of Panama and Mesoamerica: Selected Studies. Oxford University Press. Pp. 549-553. xxii + 692 pp.
LINDSEY, A. W., E. L. BELL, AND R. C. WILLIAMS, JR. 1931. The Hesperioidea of North America. Denison Univ. Bull. Jour. Sci. Lab. 26: 1-142.
Chiba, H., 2009. A revision of the subfamily Coeliadinae (Lepidoptera: Hesperiidae). Bulletin of the Kitakyushu Museum of Natural History and Human History Series A(7): 1-102.
Freeman, H.A. 1955. Four new species of Megathymus (Lepidoptera, Rhopalocera, Megathymidae). American Museum Novitiates, 1711:1-20.
Evans, 1955 A Catalogue of the American Hesperiidae, indicating the classification and nomenclature adopted in the British Museum. Part IV. Hesperiinae and Megathyminae Cat. amer. Hesp. Brit. Mus. 4: 499pp, pls. 54-88
Wooley, R. L., L. C. Keenan, M. N. Nelson and R. E. Stanford. 1991. Oviposition behavior and nectar sources of the pawnee montane skipper, Hesperia leonardus montana (Hesperiidae). Journal-of-the-Lepidopterists'-Society. 45(3):239-240.
The matrix of characters can be converted into the following 4 strings of binary characters: 0 for "-" and 1 for "+", formatted for TNT (maximum parsimony program).
xread 'morphological and behavioral characters' 80 4 Macrosoma 01000000001111000100001110111010011100111000001000100110110000??1001001100000?10 Coeliades 11100110000010010110001011000010?010001110000100000?01001010011?1111111000011001 Megathymus 10011001110000100001110100000101100011000111100011011001001111010101000010001100 Hesperia 101011011000001010010101000001101000010110110001100100001010011?0100000011100000 ;
TNT produces the following tree, which is rooted with Macrosoma:

Apparently, the tree is the same as the one obtained from DNA sequences. Significance of the tree is characterized by Bremer Support Value of 20. Warren et al. (2009) write: "We refer to support values as giving weak (BS 1–2), moderate (BS 3–5), good (BS 6–10) or strong (BS >11) support when discussing our results". Apparently, value of 20 would be off-scale strong. Bootstrap support value for this tree is 100% out of 100,000 replications.
To probe the character matrix further, we used MrBayes (Bayesian method) to make the tree. MrBayes was run with default parameters with 8 simultaneous runs of 100,000 replications each, and initial 40% of replications were discarded for the posterior probability calculation. The following tree was obtained with 100% support (posterior probability of 1).
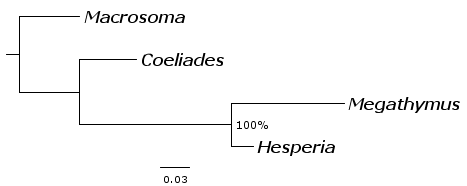
As we see the tree is again the same. Since MrBayes computes branch lengths, we notice that Megathymus branch is particularly long. This branch accounts for the characters unique to Megathymus.
Finally, do we really need all 80 characters to support this tree? To test this idea, we took the first 40 characters (from 1 to 40, this excludes genitalia, etc.):
xread 'morphological and behavioral characters' 40 4 Macrosoma 0100000000111100010000111011101001110011 Coeliades 11100110000010010110001011000010?0100011 Megathymus 1001100111000010000111010000010110001100 Hesperia 1010110110000010100101010000011010000101 ;
TNT tree turns out to be the same, albeit with smaller Bremer Support of 13, which is considered "strong" nevertheless:
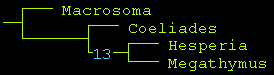

Reciprocally, we can take the remainder of the matrix (characters from 41 to 80, excluding adult morphology, except genitalia):
xread 'morphological and behavioral characters' 40 4 Macrosoma 1000001000100110110000??1001001100000?10 Coeliades 10000100000?01001010011?1111111000011001 Megathymus 0111100011011001001111010101000010001100 Hesperia 10110001100100001010011?0100000011100000 ;
TNT tree is again the same. Bremer Support is 6, which is "good":
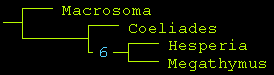

Clearly, just 1/2 of the character matrix is enough to support this phylogeny.
Conclusions:
As a last resort, we apply an unusual method of phylogeny reconstruction. We take the following images.
 |
 |
 |
 |
| 1. Macrosoma tripulata | 2. Coeliades forestan | 3. Megathymus streckeri | 4. Hesperia leonardus |
Each pixel in each image is a set of 3 numbers: {red,green,blue}, e.g. {120,32,230}. We can subtract these values between any pair of images for all pixels, square these values, average and take a square root. For instance, if a pixel in a row 45, column 176 of image 1 is {30,134,255} and a pixel in a row 45 column 176 of image 2 is {120,76,109}. Subtracting their values we get {−90,58,146}, after squaring it becomes {8100,3364,21316}, after averaging we have ~10926.7, and its square root is ~104.5. This is done for all pixels in the images and all the results are averaged. As a result, for each pair of images we have a value that characterizes distance between them. Apparently, an image compared with itself will give a distance of 0. The largest distance will be 255. This is because all-black image has all pixels {0,0,0} and all white image has all pixels {255,255,255}. After subtraction, squaring, averaging and taking a square root we get 255. For the 4 images above, the pairwise distance matrix will be as follows:
Macrosoma Coeliades Megathymus Hesperia Macrosoma 0 101.207 122.395 101.489 Coeliades 101.207 0 82.510 74.418 Megathymus 122.395 82.510 0 74.652 Hesperia 101.489 74.418 74.652 0
Here is a matrix of absolute differences of subtracted images used to compute the distances. We see that the difference between Macrosoma and Megathymus (1st row 3rd column) is the largest as this image is the lightest, and the distance between Coeliades and Hesperia (2nd row 4th column) is rather small, as this image is darker.
| 1. Macrosoma tripulata | 2. Coeliades forestan | 3. Megathymus streckeri | 4. Hesperia leonardus |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
It might be easier to see the images if their RGB values are subtracted from 255, i.e. black becomes white and white becomes black. The result is, of course, the same: the difference between Macrosoma and Megathymus (1st row 3rd column) is the largest as this image is the darkest, and the distance between Coeliades and Hesperia (2nd row 4th column) is rather small, as this image is lighter.
| 1. Macrosoma tripulata | 2. Coeliades forestan | 3. Megathymus streckeri | 4. Hesperia leonardus |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
The same images, but in black-and-white (averaging between red, green and blue).
| 1. Macrosoma tripulata | 2. Coeliades forestan | 3. Megathymus streckeri | 4. Hesperia leonardus |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
Anyway, using this method, we obtain the following distance matrix:
Macrosoma Coeliades Megathymus Hesperia Macrosoma 0 101.207 122.395 101.489 Coeliades 101.207 0 82.510 74.418 Megathymus 122.395 82.510 0 74.652 Hesperia 101.489 74.418 74.652 0
This distance matrix can be converted to a tree using Fitch-Margoliash method as implemented in Phylip "fitch" routine:
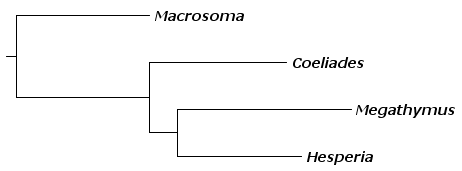
The tree has the same topology as all other trees on this page: it groups Megathymus and Hesperia as sister taxa. It also shows long Megathymus branch.
Conclusions: