
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|
|
Science is about discovery. Finding new bits of knowledge about nature is always exciting. It is even more tantalizing when the discovery conflicts with common sense and established understanding. Such discoveries should be analyzed carefully to avoid mistakes.
Analysis of DNA sequences promises to revolutionize taxonomy and to put it on a more objective basis. Within the last decade, stunning progress has been made by the Niklas Wahlberg group. The group pioneered application of major DNA techniques to Butterflies, including the polished protocol of data acquisition, and maintains wonderful web sites: The Nymphalidae Systematics Group and Nymphalidae.net. Wahlberg and colleagues obtained partial DNA sequences for many species of Nymphalidae. Many interesting and unexpected discoveries came to light as a result of this effort. For instance, it was found that Charaxes belongs to Satyrine clade, and perhaps even more surprisingly, Limenitis belongs to Heliconiine clade.
Extraordinary claims require extraordinary evidence. Here we apply various tree-building methods to available DNA sequences with the goal to convince ourselves about the validity of these clades within Nymphalidae.
Methods: All sequences used in this study were retrieved from the GenBank database as reported by Wahlberg et al. (2005). Sequences were aligned using the MUSCLE server at EBI. The trees were reconstructed using 3 phylogenetic methods (PhyML, BioNJ and MrBayes) as implemented by the phylogeny.fr server with default parameters, and the 4th program TNT, which was downloaded and executed locally. Trees were visualized with TreeView and ATV. Tutorial about how to perform these procedure is available from here.
Data: Partial DNA sequences of two nuclear (Elongation factor-1 α [EF1-a], wingless [wg]) and one mitochondrial (cytochrome c oxidase subunit I [COI]) genes from 19 Nymphalidae species, their concatenated alignment and separate alignments (EF1, COI, WG), and trees built by TNT, BioNJ, PhyML, MrBayes can be downloaded as text files using the links in this sentence.
Discussion: The main results is that major groups inside Nymphalidae stand as monophyletic regardless of the gene analyzed and the method of phylogenetic analysis. Currently, there are 4 major classes of methods to reconstruct trees. These methods are based on quite different principles, except that the general idea of Occam's razor (minimize the number of probable DNA changes) in applied everywhere. It is hardly possible to imagine a phylogenetic method that is not based on Occam's razor.
Maximum parsimony, enumerates differences between sequences and explicitly finds the tree that can be explained by the smallest number of changes, where each difference is a single change and no difference is no change. Distance-based methods estimate expected number of changes between each pair of sequences treating the process of changes as some sort of radioactive decay, i.e. the absence of difference does not mean the absence of change (changes can be followed by a back-substitution to the original state), and a difference may represent multiple changes at this position. Algebraic transformations of these distances result in a tree with certain topology. Maximum likelihood methods attempt to explicitly model substitution process and find a tree that gives the highest probability to the observed data. Bayesian methods, while quite related to maximum likelihood and use similar models to describe substitution process, do the opposite and find the tree, the observed data give the highest probability to. In any case, these 4 types of methods represent all that is available for us to use. We've chosen one software implementation per method: TNT for maximum parsimony, BioNJ for distance, PhyML for maximum likelihood and MrBayes for Bayesian methods. These methods are freely available either for download (TNT, MrBayes) or as servers.
Trees shown here are unrooted, as only Nymphalidae sequences were used (no outgroup). However, since there is good evidence that Libythea is a basal group, one can look at the trees as being rooted with Libythea. First, we show the trees from combined data of all 3 genes (two nuclear and one mitochondrial):

A tree of Nymphalidae built using maximum parsimony method (program
TNT). Numbers by the branches indicate Bremer support values.

A tree of Nymphalidae built using maximum likelihood method PhyML
and displayed with TreeView.
Numbers by the branches indicate probability.
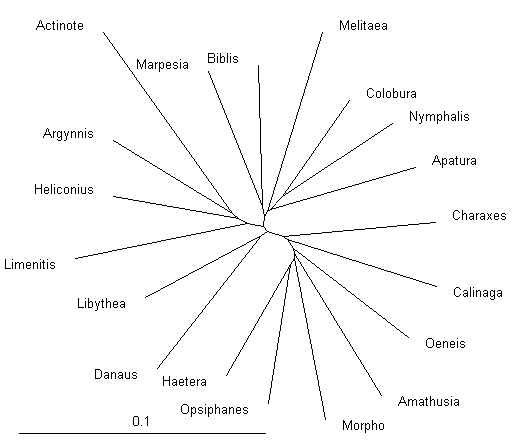
A tree of Nymphalidae built using distance-based method BioNJ
and displayed with TreeView.
Numbers by the branches indicate bootstrap support.

A tree of Nymphalidae built using Bayesian method MrBayes
and displayed with TreeView.
Numbers by the branches indicate probability.
Comparing the trees with each other we see that while there are some differences between them, for instance, Morpho is joint with Calinaga in parsimony (TNT) tree and Morpho is closer to Opsiphanes in all other trees, however, the main clades are the same. We see that Charaxes, Calinaga, Oeneis, Haetera, Amathusia, Morpho and Opsiphanes form a clade (are all together as a subtree that holds on a single branch) in all 4 trees. Another such stable clade is Limenitis, Heliconius, Actinote and Argynnis. Apparently, we are able to reproduce the results of Wahlberg et al. (2005), who used only Parsimony and Bayesian methods. We see that Charaxes groups with Satyrines and Limenitis groups with Heliconiines. This grouping is robust to the method used – all four classes of methods gave the same grouping. As illustrated here, different methods can yield different results, so it is not always that groupings generated by these methods are the same. Looking at the trees above, we don't see any unusually long branches (except Actinote), so it is unlikely that Long branch attraction was a cause for positioning of Charaxes and Limenitis.
However, is this grouping statistically significant as judged by each method? Bremer support values are shown in the parsimony tree above. Three remaining trees are displayed below in a different way. The trees are rooted with Libythea. Values signifying statistical support are shown in red by the branches:
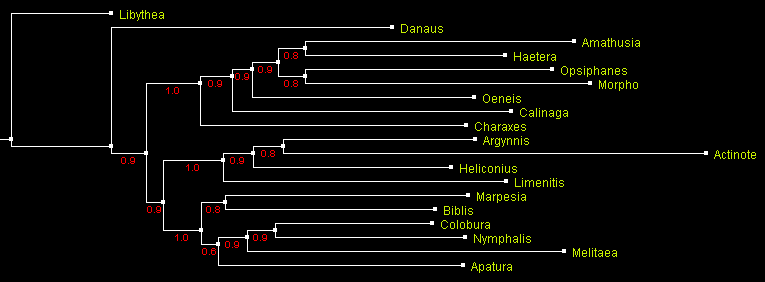
A tree of Nymphalidae built using maximum likelihood method PhyML
and displayed with ATV.
Numbers by the branches indicate probability.
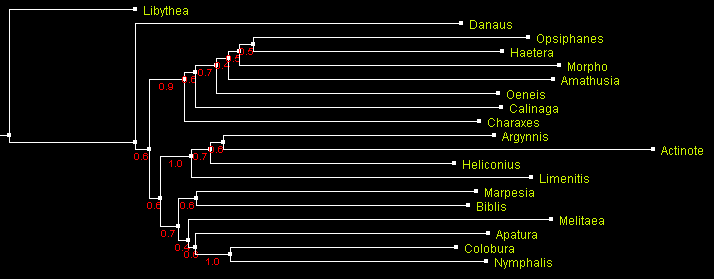
A tree of Nymphalidae built using distance-based method BioNJ
and displayed with ATV.
Numbers by the branches indicate bootstrap support.

A tree of Nymphalidae built using Bayesian method MrBayes
and displayed with ATV.
Numbers by the branches indicate probability.
Apparently, these two clades received ~100% support in all but one (~90%) cases and are among the first most strongly supported clades. What are the reasons to doubt them? Finally, DNA sequences for each gene were analyzed separately, to check whether combination of these sequences in one dataset was causing an artifact. These 3 · 4 = 12 trees are shown below.
Nuclear gene Elongation factor-1 α [EF1-a] trees.
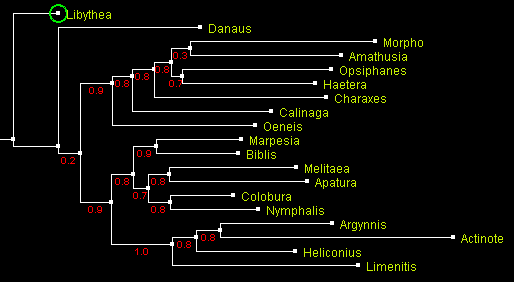
A tree of Nymphalidae built using maximum likelihood method PhyML
and displayed with ATV.
Numbers by the branches indicate probability.

A tree of Nymphalidae built using distance-based method BioNJ
and displayed with ATV.
Numbers by the branches indicate bootstrap support.
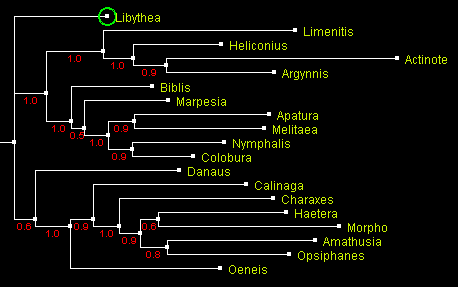
A tree of Nymphalidae built using Bayesian method MrBayes
and displayed with ATV.
Numbers by the branches indicate probability.

A tree of Nymphalidae built using maximum parsimony method (program
TNT). Numbers by the branches indicate Bremer support values.
Nuclear gene wingless [wg] trees.

A tree of Nymphalidae built using maximum likelihood method PhyML
and displayed with ATV.
Numbers by the branches indicate probability.
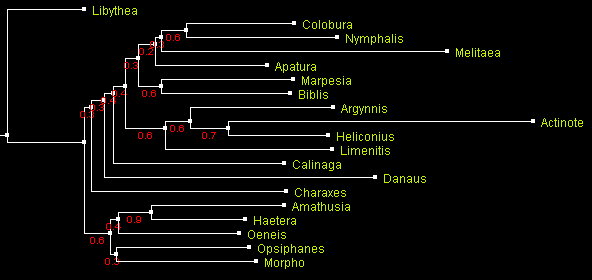
A tree of Nymphalidae built using distance-based method BioNJ
and displayed with ATV.
Numbers by the branches indicate bootstrap support.
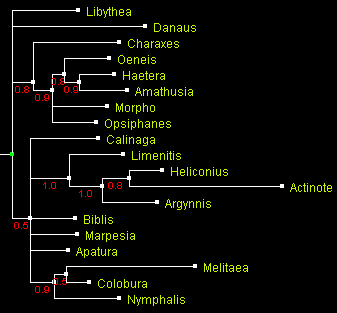
A tree of Nymphalidae built using Bayesian method MrBayes
and displayed with ATV.
Numbers by the branches indicate probability.

A tree of Nymphalidae built using maximum parsimony method (program
TNT). Numbers by the branches indicate Bremer support values.
mitochondrial gene (cytochrome c oxidase subunit I [COI]) trees.

A tree of Nymphalidae built using maximum likelihood method PhyML
and displayed with ATV.
Numbers by the branches indicate probability.

A tree of Nymphalidae built using distance-based method BioNJ
and displayed with ATV.
Numbers by the branches indicate bootstrap support.
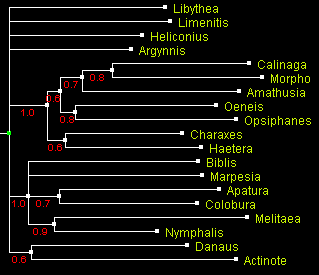
A tree of Nymphalidae built using Bayesian method MrBayes
and displayed with ATV.
Numbers by the branches indicate probability.

A tree of Nymphalidae built using maximum parsimony method (program
TNT). Numbers by the branches indicate Bremer support values.
Inspection of trees shows that in all strongly supported clades (>0.9) Charaxes is grouped with Satyrines and Limenitis is grouped with Heliconiines. If the statistical support is weak, it means sequences are not long enough (not enough data) to derive conclusions.
Conclusions: